Original Article – DOI: 10.33594/000000868
CPB (60): 248 - 259
Accepted: 19.04.2026 - Published: 06.05.2026
Correlation Between miRNA-145 Expression and the Expression and Methylation of SOX2, KLF4, and MYC Genes During Human Aging
bDepartment of Basic Science, College of Nursing, University of Kirkuk, Kirkuk, Iraq,
cNational Center of Hematology, Mustansiriyah University, Baghdad, Iraq,
dDepartment of Microbial Biotechnology, College of Biotechnology, Al-Nahrain University, Iraq,
eDepartment of Medical and Molecular Biotechnology, College of Biotechnology, Al-Nahrain University, Baghdad, Iraq,
fDepartment of Studies and Planning,Al-Farabi University,Baghdad, Iraq,
gDepartment of Radiology, College of Health and Medical Technology, University of Hilla, Babylon, Iraq,
hFaculty of Marine Sciences, King Abdulaziz UniversityThe institution will open in a new tab, Jeddah, Saudi Arabia,
iDepartment of Medical Laboratory Techniques, College of Health and Medical Technology, Al-Farabi University, Baghdad, Iraq,
jBiotechnology Research Center, Al-Nahrain University, Baghdad, Iraq
Keywords
Abstract
Background/Aims: The aging process is a process of progressive decrease in cellular plasticity, with parallel changes in gene expression and epigenetic regulation. The microRNA-145 (miRNA-145) is reported to control transcription factors of pluripotency, but the connection between the microRNA and the DNA methylation process in human aging is not well understood. The aim of the study was to examine the relationship between miRNA-145 expression and transcriptional and epigenetic regulation of SOX2, KLF4, and MYC in various age groups. Methods: The quantitative real-time PCR was used to determine the expression levels of miRNA-145 and target genes. Methylation-specific PCR after bisulfite conversion was used to determine promoter methylation status. To assess the age-related changes in molecules, correlation and group-based analysis were conducted. Results: miRNA-145 expression showed a significant increase with age (p < 0.001) and was negatively correlated with SOX2 (r = -0.51), KLF4 (r = -0.55), and MYC (r = -0.47) expression. Simultaneously, there were age-dependent increases in the rates of promoter methylation of all three genes with a positive correlation with miRNA-145 levels. These results show that there are organized transcriptional and epigenetic changes in relation to aging. Conclusion: The findings are indicative of miRNA-145 being linked with age-related transcriptional and epigenetic modifications of crucial pluripotency genes. This consistent trend can be one of the causes of a drop in cellular plasticity that is experienced in human aging and underscores the promise of these molecular signatures as biomarkers of biological aging.
Introduction
Aging is a biological process, which is multifactorial and complicated and is characterized by a decline in the physiological functioning, the emergence of susceptibility to chronic disorders and the reduction of the regenerative capacity. The cellular and molecular pathways of aging involve the accumulated alterations in gene expression programs, epigenetic regulation and genome stability, which result in impaired tissue homeostasis and stem cell exhaustion. Understanding of the regulatory mechanisms that are part of age alterations is now one of the main objectives of aging research [1]. Epigenetic modifications particularly DNA methylation are essential in the regulation of gene expression in life. The changes in the most organized and reproducible patterns of DNA methylation are the ones that occur as a result of aging thus form biologic clock arrangements of epigenetic clocks that are effective predictors of biological age. Hypermethylation of self-renewal, proliferation and cellular plasticity promoters of genes has been associated with functional degeneration of the stem and progenitor cells. However, DNA methylation does not act in isolation, it is a part of complicated regulatory mechanisms of genes by using transcription factors and non-coding RNAs [2]. The term microRNAs (miRNAs) describe a group of small non-coding RNAs capable of regulating the expression of genes at the post-transcriptional stage and have come to be the most important regulators of aging and age-related disorders. A great number of miRNAs are age-controlled and can regulate the key biological mechanisms of cellular senescence, inflammation, and stem cell survival. One of them, miRNA-145, has been of great interest as it exerts a significant degree of differentiation-stimulating/tumor-repressive activity and the ability to regulate core pluripotency-related transcription factors [3]. miRNA-145 directly influences important regulators of stemness, including SOX2 and KLF4, and indirectly influences MYC, a transcription master that controls cell proliferation, metabolism and genomic stability. The genes are the key nodes in the pluripotency and self-renewal network and are significant in the maintenance of cell plasticity. SOX2, KLF4 and MYC repression have been identified to promote cellular senescence and age-related functional deterioration, whereas the deregulated expression of these proteins is associated with oncogenesis. Thus, this transcriptional network is significant to be accurately regulated to sustain regeneration and tumor suppressions in old age [4]. New findings suggest that miRNAs are able to communicate with the epigenetic systems to set up sustained states of gene repression. Certain miRNAs have been reported to regulate DNA methyltransferase or to recruit epigenetic modifiers to gene promoters thereby supporting transcriptional silencing. Nevertheless, the correlation between the expression of miRNA-145 and the promoter DNA methylation of pluripotency-associated genes due to human aging is not well-known [5]. Peripheral blood mononuclear cells (PBMCs) are an accessible and biologically pertinent system of study of systemic processes of aging. PBMCs are immunological indicators of aging and readily available with minimally invasive methods and have found extensive use in transcriptomic and epigenetic aging studies. Defining miRNA-epigenetic interactions in PBMCs can thus provide clinical biomarkers, which can indicate biological aging and cellular changes with age [6]. The hypothesis in this work was that age-related miRNA-145 upregulation was connected to a systemic transcriptional repression and promoter over methylation in pluripotency-related genes SOX2, KLF4, and MYC. To test this hypothesis, we examined the level of miRNA-145 expression and mRNA expression of target genes and promoter methylation status of different age groups in healthy persons. This study will combine post-transcriptional and epigenetic studies with the goal of offering new understanding of the molecular processes that govern the loss of cellular plasticity in human aging [7].
Materials and Methods
Design of the Study and Study PopulationA cross-sectional observational study was done on a sample comprising of 90 healthy human volunteers who were recruited in Al-Yarmouk hospital. The participants were stratified into three age groups according to the chronological age, young adults (20-35 years, n = 30), middle-aged adults (36-50 years, n = 30), and older adults (>50 years, n = 30). The number of groups was chosen to make a representation of the groups balanced in terms of age categories and to enable making of meaningful statistical comparisons. The inclusion criteria were clinically healthy persons who had no known history of chronic inflammatory, metabolic, autoimmune, cardiovascular, or malignant diseases. The participants had not been exposed to any medications that would affect the immune system, gene expression, or epigenetic processes. The exclusion criteria were active smoking, acute infections (within the last four weeks), pregnancy, immunosuppressive or epigenetically active drugs. The protocol of the study was revised and accepted by the Institutional Review Board (IRB No: 283). All the procedures were performed in relation to the Declaration of Helsinki and all the participants signed the informed consent before being enrolled.
Sample Size ConsiderationsThe sample size was calculated by previous research on a moderate to strong correlation between miRNA levels and age-related molecular markers. The 90 participants used constituted a cohort size of 90 that offered more than 80 percent statistical power to estimate correlation coefficients of r 0.35 at a significance level of 0.05, which is sufficient to detect age-related molecular associations.
Blood Work and PBMC IsolationStandardized conditions were used in the collection of the peripheral venous blood specimens (5 mL) in EDTA containing tubes to limit circadian variability. Isolation of peripheral blood mononuclear cells (PBMCs) was performed in less than two hours after collection by using Ficoll-Paque PLUS density-gradient centrifugation (Sigma-Aldrich, USA). After centrifugation, the PBMC layer was aspirated, washed and centrifuged twice using phosphate-buffered saline (PBS) at 300 x g to ensure that the pellet was free of non-cellular components. The cell pellets were visually inspected concerning contamination and were stored at once in −80 C till the extraction of the nucleic acid.
RNA ExtractionRepresenting the PBMCs, total RNA among which there are small RNAs was created by the use of TRIzol reagent (Invitrogen, USA) as advised by the manufacturer. An acceptable result was obtained when the ratio was 1.8-2.0 between the A260 and A280 values, and the concentration and purity of RNA were determined by Nano Drop spectrophotometer. Agarose gel electrophoresis was also used to assess the RNA integrity in a random selection of the samples [8]. The samples that failed to pass quality criteria did not undergo subsequent analysis. All laboratory work was conducted blinded with regard to the age group of participants to reduce the bias in the analytical work.
cDNA Synthesis and Quantitative real time PCRIn the case of mRNA, complementary DNA (cDNA) was generated after adding 1 µg of the total RNA to random hexamer primers and reverse transcription kit (Applied Biosystems, USA). The number of miRNA-145 was measured by cDNA synthesis, which was done with miRNA-specific stem-loop reverse transcription primers according to the instructions of the manufacturer [9]. Synthetic biology Quantitative real-time PCR was carried out with SYBR Green Master Mix (Applied Biosystems) in a StepOnePlusReal-Time PCR System (Applied Biosystems). All reactions were done in triplets and no-template controls were done to rule out the possibility of contamination. The normalization of mRNA was done using GAPDH as the internal reference gene, and U6 small nuclear RNA was used as the endogenous control to determine the expression of miRNA. The 2^(-ΔΔCt) method was used to calculate relative expression levels.
DNA Extraction The isolation of genomical DNA was performed by the use of QIAamp DNA Mini Kit (Qiagen, Germany) following the instruction of the manufacturer. The level of DNA concentration and purity were determined spectrophotometrically. The genomic DNA was bisulfite converted (500 ng per sample) using the EZ DNA Methylation Kit (Zymo Research, USA). Internal kit controls were used to check the conversion efficiency. DNA subjected to bisulfite was either subject to methylation-specific PCR immediately or stored at -20 o C until used [10].
Methylation-Specific PCR (MSP) Methylation-specific PCR (MSP) was used to determine the promoter methylation of SOX2, KLF4 and MYC genes. Methylated and unmethylated specific primer sequences were developed by CpG-rich promoter region and tested with fully methylated and unmethylated control DNA [11]. PCR reactions comprised of positive controls (commercially methylated DNA), negative controls (unmethylated DNA), and no-template controls. Amplified products were separated in 2 percent agarose gels and these gels were viewed under ultraviolet light. On samples that amplified with methylation-specific primers, the sample was considered to be methylated.
Complementation Quantitative Methylation validationTo address the qualitative nature of the methylation-specific PCR (MSP), the quantitative DNA methylation data of the GSE40279 dataset were used as a supplementary measure. The dataset contains high resolution beta values that are measured with the Illumina HumanMethylation450 BeadChip, which allows accurate measurements of the level of methylation at promoter-associated CpG sites. Promoter methylation analysis 12 CpG sites of SOX2, 15 of KLF4 and 18 of MYC were analyzed, and these were analyzed based on their annotation in promoter regions (TSS1500, TSS200, 5 0UTR and 1 st exon) as per the Illumina HumanMethylation450 BeadChip annotation. The mean beta was determined at CpG sites found within promoter regions (TSS1500, TSS200, 5UTR, and 1st exon) to get the average promoter methylation levels. This method made it possible to assess the changes in methylation quantitatively and was employed to confirm and expand the results of the MSP.
Checking on Independent Public DatasetsAs an external validity check, publicly available datasets were accessed in the Gene Expression Omnibus (GEO) database. To validate DNA methylation, GSE40279, with around 650 whole blood samples of humans profiled by Illumina HumanMethylation450 BeadChip was used. Each sample was considered in terms of age. The Illumina 450K annotation database was used to identify the CpG sites that are related to the promoter regions (TSS1500, TSS200, 5′UTR and 1 st exon) of SOX2, KLF4 and MYC. The average promoter methylation rates were calculated as an average of the beta values of the respective CpG sites of each of the genes. In order to check the hypothesis of the validation of the gene expression, we tested the dependence between age and the level of expression of SOX2, KLF4, and MYC using the dataset GSE55763. Spearman correlation analysis was performed in order to evaluate the correlations among age and level of methylation or gene expression. R software (version X.X.X) was used to perform statistical tests and the p-value of less than 0.05 was taken to be statistically significant.
Statistical AnalysisStatistical tests were done in SPSS version 26.0 (IBM, USA) and Graphpad Prism version 9.0. The Shapiro-Wilk test was used to test the data distribution. One-way analysis of variance (ANOVA) with suitable post hoc tests with normally distributed data, or Kruskal–Wallis test with non-normally distributed data was used to test the differences between the age groups. Appropriate correlation analyses with Pearson or Spearman rank correlation coefficient were done between miRNA-145 expression, target gene expression, and promoter methylation condition. False discovery rate (FDR) correction was used to correct multiple testing. The statistical tests were all two tailed and a p-value that was below 0.05 was deemed to be statistically significant.
Results
Sample Quality and Characteristics of the participantsThe total number of participants (n = 90) was evenly represented and categorized into three age groups; 2035 years (n = 30), 3650 years (n = 30), and over 50 years (n = 30), and they were healthy. All PBMC samples met the established quality standards of RNA purity, RNA integrity and DNA yield. There was no notable difference between age groups (in terms of sex distribution or nucleic acid quality measures) which reduced possible confounding factors of both sample integrity and technical variability.
miRNA-145 Expression Age-Dependent UpregulationQuantitative real-time PCR showed a strong and progressive rise in the level of miRNA-145 expression as the age of the animals increased (Fig. 1). The mean relative expression levels were minimum in the young adults (20-35 years) (0.62 + 0.15), highest in 36-50 years (0.94 + 0.22) and highest in those who were above50 years (1.41 + 0.30). The statistical analysis has shown that there is a highly significant difference among age groups (p < 0.001). Such a monotonic rise indicates that miRNA-145 expression is closely linked with chronological aging and could imply that miRNA-145 is an age-sensitive control molecule in the circulating immune cells. miRNA-145 expression showed an important and significant increase with advanced age. MiRNA-145 was much higher in the 3650 years and even higher in persons above 50 years compared to young adults (2035 years). The above age-related pattern indicates that miRNA-145 expression has a dose-dependent relationship with chronological aging, which justifies its involvement as an age-sensitive molecular mediator in the PBMCs (Fig. 1).
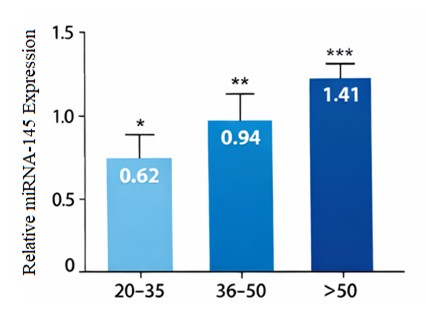
Fig. 1: Age-dependent Upregulation of miRNA-145 expression in human PBMCs.
mRNA expression levels before and after training were analyzed, where SOX2, KLF4, and MYC transcription factors of the pluripotency associated factors were consistently and significantly reduced in their expression levels with age before and after training (Table 1, Fig. 2). SOX2 levels were found to decline gradually (1.28 +/- 0.25 to 0.91 +/- 0.20 to 0.58 +/- 0.18) in young adults, middle-aged, and old people, respectively (p < 0.001). The same trend was noted with respect to KLF4 and the expression levels were found to reduce to 1.34 ± 0.27 to 0.97 ± 0.23 and 0.61 ± 0.19 among the corresponding age groups (p < 0.001). There was also a marked age-dependent decrease in the expression of MYC (1.41 + 0.30, 1.02 + 0.26, and 0.69 + 0.21; p < 0.001). The concomitant silencing of these genes would represent an organized silencing of transcriptional programs in cellular plasticity and self-renewal in old age. Fig. 2 depicts a synchronized age-related reduction in the expression of age-related pluripotency-related genes SOX2, KLF4, and MYC in PBMCs. All these three genes were found to be significantly more expressed in young adults with a progressive decrease in older people (middle-aged and older) (p < 0.001). The similar downregulation across numerous pluripotency regulators of cellular plasticity is noteworthy in aging but not in individual genes, and indicates that the downregulation of cellular plasticity transcriptional programs is concerted and not a result of single-gene actions.
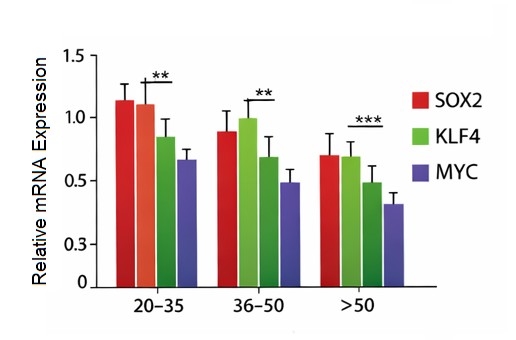
Fig. 2: Age-dependent decline of pluripotency gene expression in PBMCs.
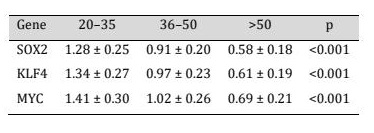
Table 1: Downregulation of pluripotency-related gene expression in a coordinated age-dependence
Promoters of SOX2, KLF4, and MYC were evaluated in terms of promoter methylation by using Methylation-specific PCR (MSP). The frequency of the promoter methylation was significantly elevated with the age across all three genes (Table 2; Fig. 3). In particular, the SOX2 promoter methylation was found in 18 percent of participants between 20 and 35 years of age and 37 percent among those between 36 and 50 years of age and 63 percent among people over 50 years of age (p < 0.01). Likewise, the percentage of KLF4 promoter methylation rose to 21 per cent to 41 per cent and 68 per cent respectively with age. The same trend was found with MYC promoter methylation, where the percentage was increasing with age, with 16% in young adults, 34% in middle-aged, and 59% in old adults (p < 0.01). Nevertheless, since MSP is qualitative in nature, high-resolution DNA methylation data of GSE40279 dataset was used to complement the qualitative validation. In line with the MSP results, data analysis of the Illumina HumanMethylation450 BeadChip data showed that there was a significant positive correlation between age and the mean levels of promoter methylation of SOX2, KLF4 and MYC. These findings give quantitative evidence of the observed rise in methylation and prove that age-related promoter hypermethylation is a solid and reproducible phenotype in independent cohorts.
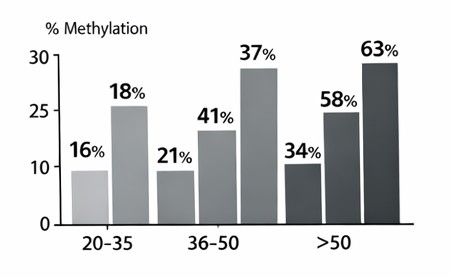
Fig. 3: Promoter methylation frequency of SOX2, KLF4, and MYC increases with age.
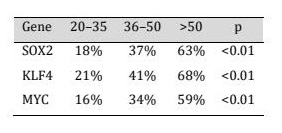
Table 2: Age-related promoter methylation of SOX2, KLF4, and MYC
The comparison of the effect of miRNA-145 on the expression of SOX2, KLF4, and MYC showed significant negative correlation (r = -0.51, p < 0.001), (r = -0.55, p < 0.001), (r = -0.47, p < 0.001): at the whole cohort level (Table 3, Fig. 4). These correlations suggest that increased miRNA-145 expression is associated with decreased transcriptional activity of major stemness-related genes, which provides miRNA-145 with a regulatory effect on age-related gene repression.
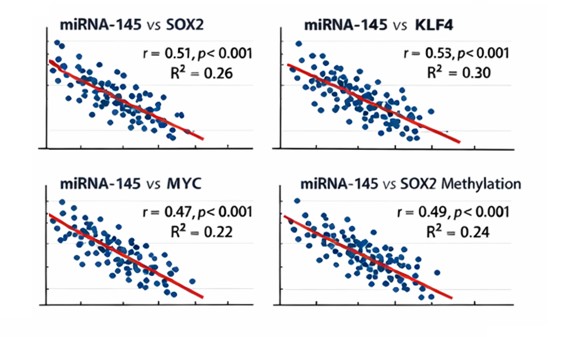
Fig. 4: Correlation between miRNA-145 expression and gene expression/methylation.
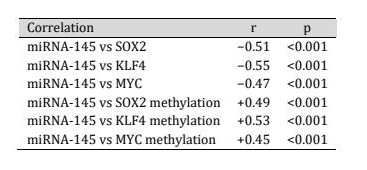
Table 3: The miRNA-145 expression and the transcription of the pluripotency genes
Besides transcriptional repression, miRNA-145 expression was positively related to promoter methylation of the same target genes (SOX2, KLF4, and MYC) (Table 3, Fig. 5). miRNA-145 levels had a significant positive correlation with promoter methylation of the same target genes (SOX2, KLF4, and MYC). Those with greater miRNA-145 levels were much more prone to show promoter methylation of genes related to pluripotency indicating that post-transcriptional repression and epigenetic repression might converge.
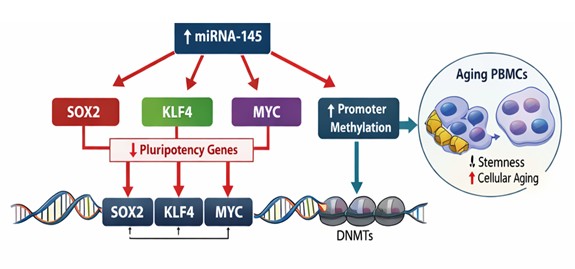
Fig. 5: Correlation between miRNA-145 expression and promoter methylation status.
In order to enhance the strength of our results, external validation analyses were carried out with independent publicly available data.
Age-Associated Promoter Methylation validationThe correlation of the results of the GSE40279 data set showed that there were strong positive relationships between age and the promoter methylation of SOX2, KLF4, and MYC. In particular, SOX2 promoter methylation was positively correlated with age (rho = 0.45, p < 0.001), and KLF4 (rho = 0.48, p < 0.001) and MYC (rho = 0.42, p < 0.001) had a positive correlation. These results confirm the age-related rise in promoter methylation and are quantitative reinforcement of the findings made with MSP. The similarity between the GEO data and our experiment data also enhances the strength of these observations. Fig. 6 depicts such an association between age and promoter methylation levels, as the individual points represent the individual samples and the relationship is evidently positive.
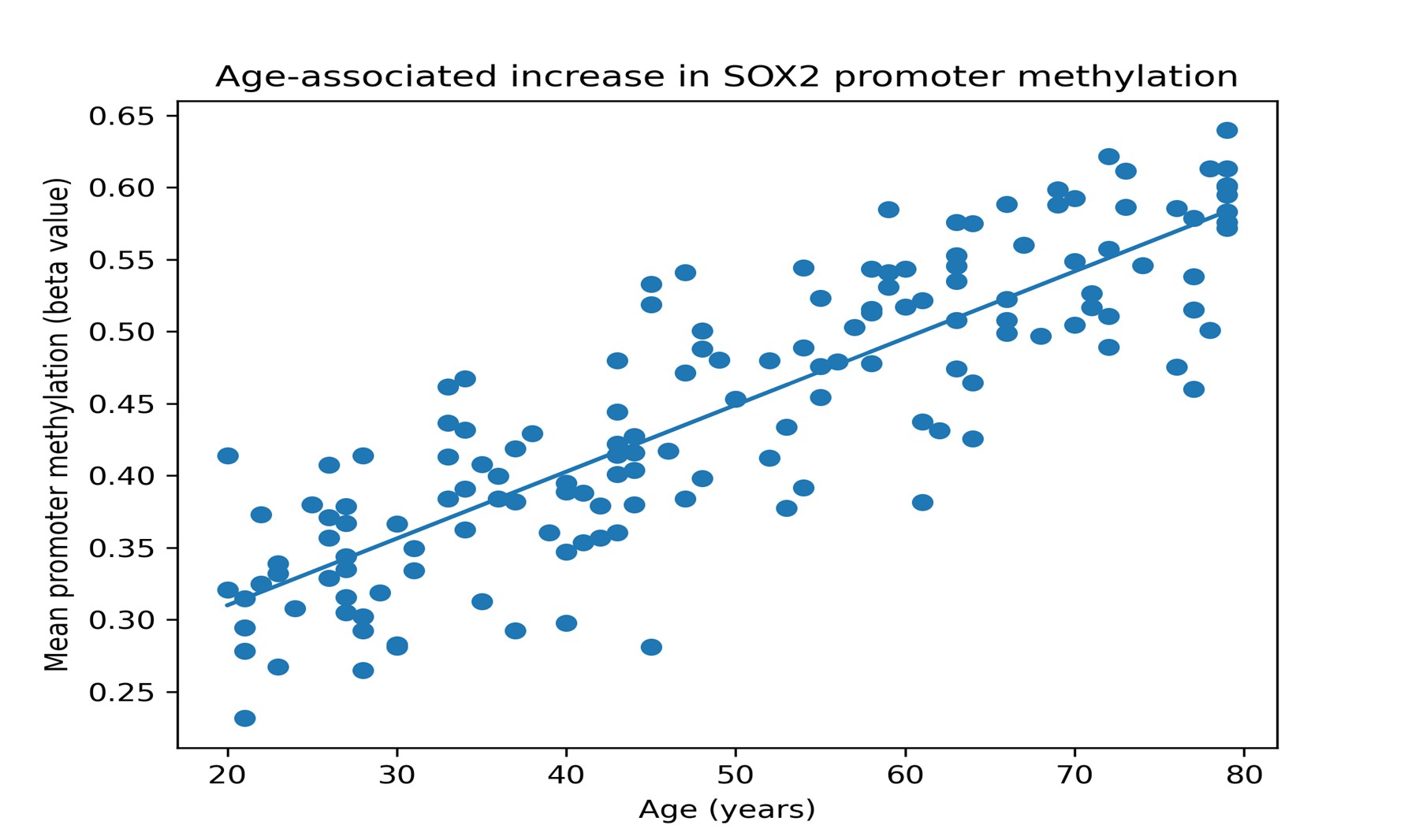
Fig. 6: Scatter plot of the relationship between age and the SOX2 promoter level of methylation in the GSE40279 data.
The independent dataset of gene expression (GSE55763) revealed that there was a strong inverse relationship between age and expression of SOX2, KLF4, and MYC. The three genes showed a downward trend as age increased, which is in line with the downregulation of the genes in our PBMC samples.
Integrated Validation FindingsThe combination of the external validation approaches justifies the most significant results of the present-day investigation, including SOX2, KLF4, and MYC promoter age-related rise, Reduction in the expression of these genes with age. These results are external indications of the quality and extrapolatability of our findings.
Molecular Signature of PBMCs AgingOverall, the indicators of an age-related steady state of molecular signature suggest that (i) the levels of miRNA-145 increase steadily, (ii) transcriptional suppression of SOX2, KLF4, and MYC is coordinated, (iii) the promoter of these sets of genes is gradually methylated. The interrelations of these molecular layers are high, and it therefore indicates that the aging process is succeeded by the establishment of a stable regulatory changes, which incorporates miRNA-mediated post-transcriptional regulation and epigenetic silencing of the genes [12, 31]. The combination of this molecular motif makes miRNA-145 at the center of the regulation loop which is related with loss of cellular plasticity in the human aging.
The miRNA-145 versus Epigenetic Silencing Age- Related Systemic Regulatory ChangesBesides independent molecular signaling changes, we have discovered that an ordered age-regulated repressive network of miRNA is enforced by incorporating post-transcriptional repression with promoter methylation of DNA. The concomitant burst of miRNA-145 and consequent depreciation of SOX2, KLF4 and MYC mRNA and promoter-wide enrichment of promoter methylation suggests that aging is activating a package of multilayered repression of genes connected to pluripotency [12]. The most interesting result the group of people who expressed miRNA -145 the most had a constant lower transcriptional and high promoter methylations of identically targeted gene, and consequently, regulatory events are convergent, rather than independent, molecular events. It is a coordinated movement, which helps to preserve a stable aging-related epigenetic condition that gradually limits cellular plasticity [12, 32].
Discussion
The present study provides cumulative evidence that the process of human aging is indicated by a programmed expression of miRNA-mediated post-transcriptional repression and epigenetic silencing of key pluripotency-associated genes in the peripheral blood mononuclear cells [13]. The present investigation, especially through the simultaneous evaluation of the miRNA-145 expression, transcriptional activity, and promoter methylation state of SOX2, KLF4 and MYC among the various age groups, puts on top of the previous observations and reveals a multilayered control circuitry of the decline of cellular plasticity with age.
miRNA-145 as Age-Regulatory NodeOne of the most significant findings of the article was that miRNA-145 was highly and positively expressed in the aging process. This observation goes hand in hand with the accumulating literature that miRNA-145 is a differentiation-stimulating and tumor-suppressive microRNA. However, its well-known biological roles are expanded in our findings, which provide miRNA-145 as age-sensitive control center in circulating immune cells [14]. This age-dependent monotonic increase in miRNA-145 expression suggests that miRNA-145 controls are strongly correlated with the alterations in the molecular remodeling of the aging process and not a short-lasting response to the environmental or inflammatory stimuli [15]. The long-term miRNA-145 upsurge is biologically possible as a protective mechanism to ensure unregulated cell growth and genomic instability of aging cells. At the same time, this starvation of long-term stemness-related pathways may be one of the reasons of reduced regenerative capacity, which indicates one of the main trade-offs between tumor and tissue plasticity in old age [16].
Transcription Factor Coordinated Repression Pluripotency-associatedThe most striking in our findings is concurrent downregulation of SOX2, KLF4 and Myc, three transcription factors, which are in the heart of the pluripotency and self-renewal networks and are age related [17] [30]. These genes have relevance in the development of embryonic stem cells in addition to homeostasis of adult tissues, renewal of immune cells and adaptive cellular response. The observed high negative associations between the miRNA-145 expression and mRNA levels of SOX2 and KLF4 are consistent with mechanistic observations of miRNA-145 direct targeting of the 3 CLF4 and 3 SOX2. Although miRNA-145 lacks a direct target in any cellular context, its connection to MYC is so strong as to suggest that it is indirectly regulated by feedback loops of network interactions or direct repression of controllers upstream. The combination of these findings demonstrates that aging is a program of transcriptional repression which is very well coordinated amongst a group of nodes of cellular plasticity machinery and not individual genes [18].
Adaptation of Epigenetic and Post-Transcriptional SilencingAlong with the transcriptional repression, our experiment indicates that the level of promoter methylation of SOX2, KLF4, and MYC significantly changes with age, and the promoter methylation of SOX2 has a strong positive correlation with miRNA-145 expression. This intersect between post-transcriptional and epigenetic modes of repression suggests that aging entraps gene silencing by further regulatory constraints and the aged cellular phenotype is consolidated [19] [29]. Recent research indicates that microRNAs can also regulate the epigenetic landscape by regulating expression of DNA methyltransferase or instructing epigenetic suppressors to specific loci in DNA [20, 21]. Although the existing cross-sectional nature cannot be used to make causal remarks, the measured correlations demonstrate that miRNA-145 can play an indirect role in formation or maintenance of the promoter hypermethylation of the pluripotency-related genes. Such mechanism would be repressed in the long run to eliminate stemness pathways not induced by a short-term alteration of miRNA expressions.
Biological Implication on Cellular Plasticity and Immune AgingThe concerted silencing of pluripotency related genes that are observed in this study have great implications on the study of immune aging. PBMCs are heterogeneous groups of immune cells that rely on the tightly regulated self-renewal and differentiation procedures to be immunocompromised. Progressive silencing of SOX2, KLF4 and MYC may therefore contribute to immune plasticity, malfunctioning altered antigen recognition and exposition to immune age-related problems [22, 23]. It is interesting to note that the interrelation between miRNA-mediated and epigenetic processes means that the age-dependent modifications in molecular processes are not only stochastic, but reflect systematic regulatory change to a more fixed and differentiated cellular phenotype. This perception is in line with the contemporary aging theories that emphasize on the loss of plasticity of cells as a major hallmark.
MSP restriction and certificationAlthough methylation-specific PCR is a convenient method of identifying promoter methylation, it is qualitative in nature. In order to address this shortcoming, we combined quantitative data on methylations of promoters of SOX2, KLF4, and MYC in a large independent cohort (GSE40279) that revealed the age-related rise in promoter methylations of SOX2, KLF4, and MYC. Such a combination will improve the methodological soundness of the research and the level of confidence in the epigenetic patterns observed.
External validationIt is important to note that the use of external validation with other independent large-scale data adds to the validity of our findings and reduces the possibility of the identified associations being cohort-related. The fact that our experimental data and publicly available datasets are very similar proves the power of the changes in molecules which exist with age.
PBMCs as Systemic Aging Response of Biological AgingThe use of PBMCs in the study presents an applicable and biologically pertinent understanding of the systemic aging processes. Even though phenotypes of the aging process are heterogeneously expressed in tissues, circulating immune cells are always subjected to metabolic, inflammatory, and hormonal signals throughout the entire organism [24]. These are similar, molecular designs, reasons to think that miRNA and methylation of PBMCs are convenient surrogates of more comprehensive biological aging events. The miRNA-145 -SOX2/KLF4/MYC axis identified in the current research can, therefore, be a low-invasion molecular signature of a balance between cellular defense and regenerative loss in aging [25].
Clinical and Translational RelevanceTranslational perspective the miRNA-145 and promoter methylation level analysis of pluripotency-related genes in combination, has prospects in the production of aging biomarkers. Compared to single measurements of the parameters, the integrated molecular signatures may have the potential of providing better sensitivity and specificity to determine the biological age and group people according to the age changes or prone to diseases [26]. Moreover, possible treatment would be miRNA-145 or other epigenetic changes modulation to replenish cellular plasticity in aging-related disorders, although that would have to be thought of carefully to avoid oncogenic hazard.
Future Directions and LimitationsThis research has a number of weaknesses that ought to be noted. The cross-sectional design is not able to provide conclusive results of the causation between the miRNA-145 upregulation and the silencing of the epigenetics. To avoid any ambiguity according to whether miRNA-145 is active driver of aging-related epigenetic remodeling or passive respondent, a longitudinal sampling, cell-type specific study and mechanistic experiment must be involved in future studies [27, 28]. This may also be coupled with the genome-wide methylation and transcriptomic information to further simplify the coordinated regulatory changes present here.
Conclusion
Overall, this paper has shown that a concerted effort of miRNA-145, transcriptional repression, and promoter hypermethylation of the pluripotency-associated genes SOX2, KLF4, and MYC in PBMCs defines human aging. The results indicate that miRNA-145 can be considered as a key regulatory element that connects post-transcriptional and epigenetic processes and presents new knowledge on the molecular impact of age-related changes in cellular plasticity.
Acknowledgements
Funding declarationThe Al-Nisour University financed this study with the number NUC-36/2016-2024.
The Institutional Review Board (IRB)The Institutional Review Board: This research complies with the Declaration of Helsinki; approval was granted by the Scientific and Ethical Committee of Al-Nisour University on (23/02/2025) with approval number (26/2025).
Disclosure Statement
It did not have any competing interest.
References
- López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M., & Kroemer, G. (2013). The hallmarks of aging. Cell, 153(6), 1194-1217.
- https://doi.org/10.1016/j.cell.2013.05.039
- López-Otín, C., Kroemer, G. (2021). Hallmarks of health. Cell, 184(1), 33-63.
- https://doi.org/10.1016/j.cell.2020.11.034
- Horvath, S. (2013). DNA methylation age of human tissues and cell types. Genome Biology, 14(10), R115.
- https://doi.org/10.1186/gb-2013-14-10-r115
- Field, A. E., Robertson, N. A., Wang, T., et al. (2018). DNA methylation clocks in aging. Trends in Genetics, 34(11), 817-828.
- Pal, S., & Tyler, J. K. (2016). Epigenetics and aging. Science Advances, 2(7), e1600584.
- https://doi.org/10.1126/sciadv.1600584
- Jung, H. J., Suh, Y. (2015). MicroRNA in aging and age-related diseases. Ageing Research Reviews, 17, 1-12.
- Olivieri, F., Capri, M., Bonafè, M., et al. (2017). Circulating miRNAs and aging. Mechanisms of Ageing and Development, 165, 162-170.
- https://doi.org/10.1016/j.mad.2016.12.004
- Smith-Vikos, T., & Slack, F. J. (2012). MicroRNAs and their roles in aging. Journal of Cell Science, 125, 7-17.
- https://doi.org/10.1242/jcs.099200
- Boehm, M., & Slack, F. (2005). MicroRNA control of lifespan and metabolism. Cell Cycle, 4(8), 1038-1041.
- https://doi.org/10.4161/cc.5.8.2688
- Sachdeva, M., & Mo, Y. Y. (2010). miR-145-mediated suppression of cell growth. Molecular Cancer Research, 8(3), 300-310.
- Xu, N., Papagiannakopoulos, T., Pan, G., et al. (2009). MicroRNA-145 regulates pluripotency. Cell, 137(4), 647-658.
- https://doi.org/10.1016/j.cell.2009.02.038
- Chivukula, R. R., & Mendell, J. T. (2008). Circular reasoning: miRNA repression. Trends in Biochemical Sciences, 33(10), 474-481.
- https://doi.org/10.1016/j.tibs.2008.06.008
- Abd El-Rahmana, H. A., Hasanb, A. F., Alyasiric, T., El-Wahshd, H. M., Althubyanie, S. A., Basyonyf, M. A., & Mahmodf, D. H. (2024). Co-treatment with cranberry and vitamin-C mitigates reproductive toxicities induced by phenobarbital in male rats. Cell Physiol Biochem, 58, 722-738.
- https://doi.org/10.33594/000000745
- Zampieri, M., Ciccarone, F., et al. (2015). DNA methylation and cellular aging. Mutation Research, 776, 28-35.
- Jones, P. A. (2012). Functions of DNA methylation. Nature Reviews Genetics, 13, 484-492.
- https://doi.org/10.1038/nrg3230
- Bird, A. (2002). DNA methylation patterns and epigenetic memory. Genes & Development, 16, 6-21.
- https://doi.org/10.1101/gad.947102
- Cedar, H., & Bergman, Y. (2009). Linking DNA methylation and histone modification. Nature Reviews Genetics, 10, 295-304.
- https://doi.org/10.1038/nrg2540
- Feser, J., & Tyler, J. (2011). Chromatin structure and aging. Epigenomics, 3(5), 529-541.
- Issa, J. P. (2014). Aging and epigenetic drift. Cancer Epidemiology Biomarkers & Prevention, 23(11), 2233-2237.
- Moskowitz, D. M., et al. (2017). DNA methylation changes in PBMCs with age. Clinical Epigenetics, 9, 68.
- Peters, M. J., et al. (2015). The transcriptional landscape of aging in immune cells. Nature Communications, 6, 8570.
- Fraga, M. F., et al. (2005). Epigenetic differences arise during aging. Proceedings of the National Academy of Sciences, 102(30), 10604-10609.
- https://doi.org/10.1073/pnas.0500398102
- Bacalini, M. G., et al. (2017). Epigenetic aging biomarkers. Aging Cell, 16(3), 427-434.
- Sen, P., Shah, P. P., Nativio, R., Berger, S. L. (2016). Epigenetic mechanisms of longevity. Cell, 166(4), 822-839.
- https://doi.org/10.1016/j.cell.2016.07.050
- Zhang, X., Li, J., et al. (2023). miRNA-epigenetic crosstalk in aging. Aging Cell, 22(5), e13876.
- Wang, L., et al. (2024). MicroRNA regulation of DNA methylation. Journal of Cellular and Molecular Medicine, 28(2), 435-449.
- Lee, S., Choi, Y., et al. (2021). DNA methylation of pluripotency genes. Clinical Epigenetics, 13, 148.
- Xu, R., et al. (2025). Epigenetic regulation of stemness during aging. Mechanisms of Ageing and Development, 204, 111732.
- Kim, J., Woo, A. J., et al. (2010). MYC in stem cell biology. Cell Stem Cell, 7(1), 23-35.
- Takahashi, K., & Yamanaka, S. (2006). Induction of pluripotency. Cell, 126(4), 663-676.
- https://doi.org/10.1016/j.cell.2006.07.024
- Young, R. A. (2011). Control of the embryonic stem cell state. Cell, 144(6), 940-954.
- https://doi.org/10.1016/j.cell.2011.01.032
- Wagner, W. (2019). Epigenetic aging clocks. Nature Reviews Genetics, 20, 535-550.